-
Integrated Drug Discovery
Integrated Drug Discovery
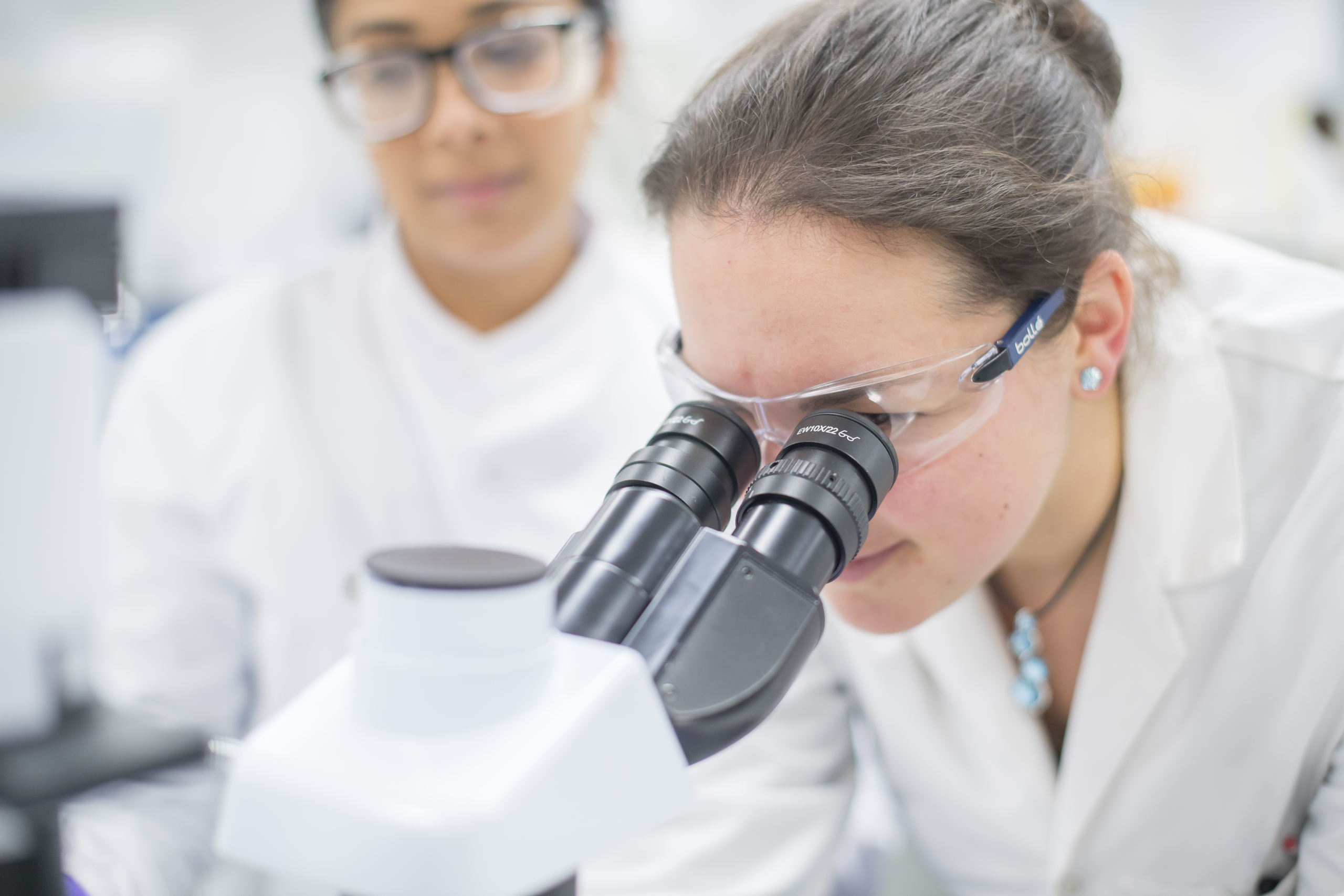
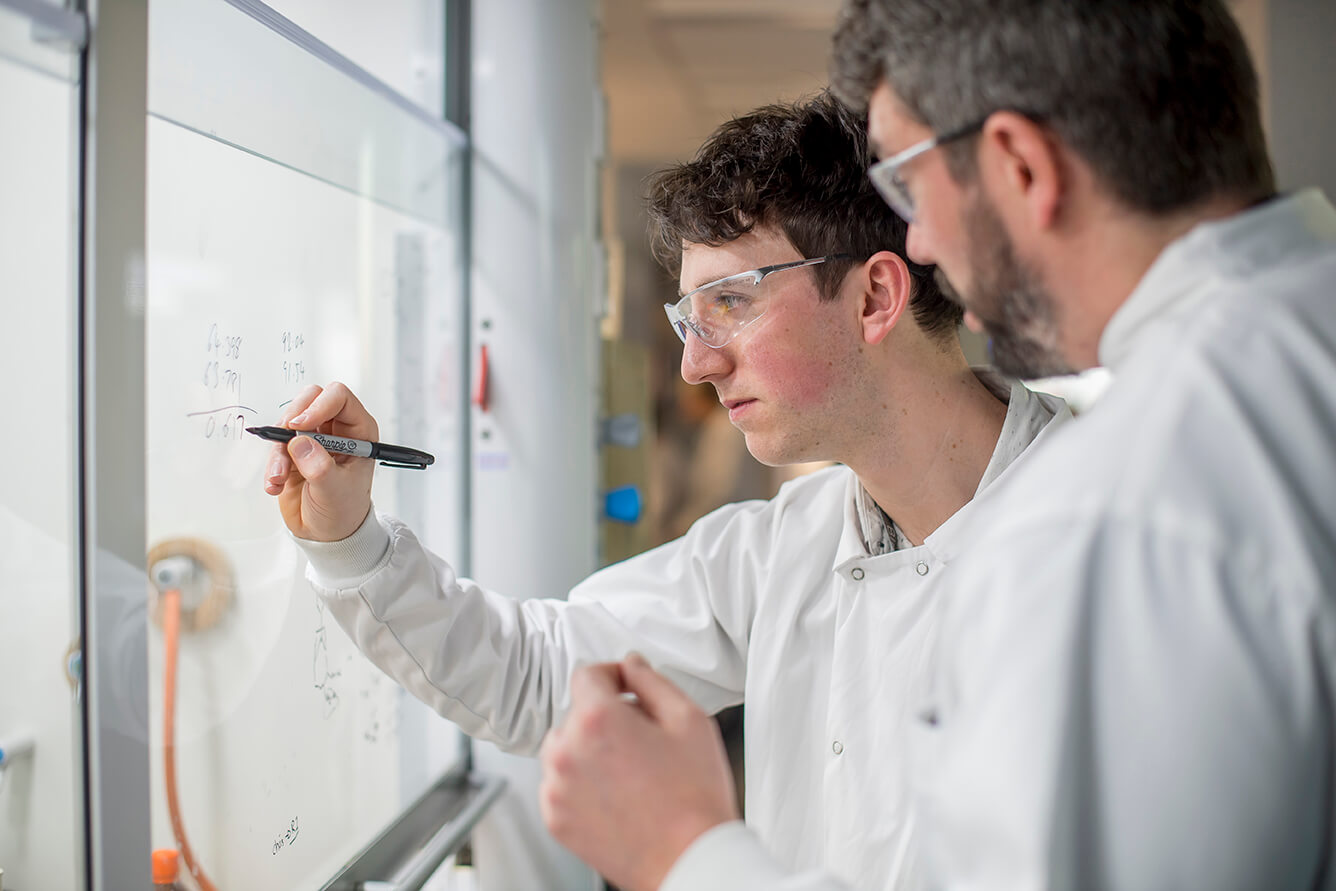
OverviewWe deliver truly integrated drug discovery programs. Our co-located teams accelerate compounds from target ID to candidate.
-
Target Identification & Validation
Target Identification & Validation
It’s important to lay strong foundations for successful drug discovery at this first stage of the process. Our integrated target identification and validation platform combines AI with expert insights, and rigorous lab validation to guide targets through robust evaluation, ready for hit discovery.
-
Hit Identification
Hit Identification
Validated, high-quality hits, delivered through integrated technologies and expert collaboration, give you a confident starting point for faster drug discovery.
-
Hit-to-Lead
Hit-to-Lead
High-quality hits with early proof of concept are de-risked and profiled, ready to accelerate your program towards lead optimization.
-
Lead Optimization
Lead Optimization
Turning promising leads into clinical candidates with speed, precision, and the scientific expertise to generate high-quality data and deliver real patient impact.
Featured Resources
-
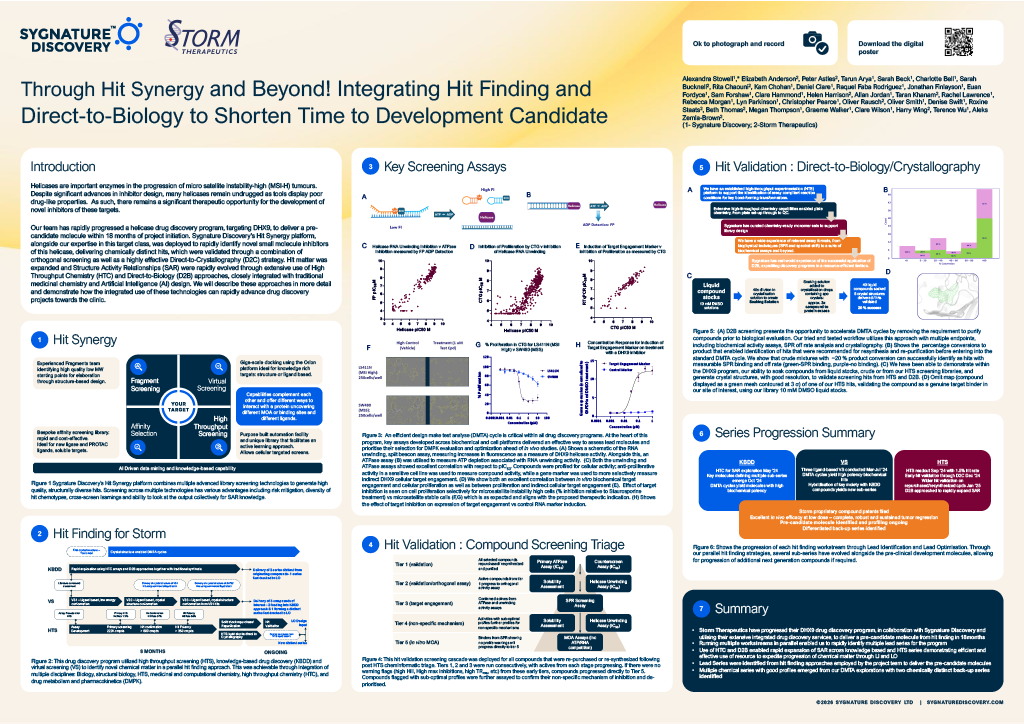
STORM Therapeutics: From Lead to Pre-Candidate Nomination in 18 Months

STORM Therapeutics: From Lead to Pre-Candidate Nomination in 18 Months
Case study