Mechanism of Action (MOA)
Understanding Mechanism of Action (MOA): Approaches, Techniques, and Characterisation
Determining the Mechanism of Action by which a compound modulates its target’s function or the biology of cells of interest is key to delivering the right compound with the right pharmacology. Sygnature Discovery is expert at applying different, complementary approaches that can be taken to identify compound Mechanism of Action. Classical enzymology techniques can be used to determine whether compounds fall into the four main classes of activity inhibition: competitive, non-competitive, mixed and uncompetitive. Furthermore, reversibility can be assessed kinetically and by mass spectroscopy, with tight binding being identified by protein ratio experiments.
Compounds of Mechanism of Action
To determine a compounds Mechanism of Action on receptor targets, classical pharmacology experiments can be used to characterise receptor agonists and antagonists, with Biophysical methods being used for the characterisation of binding kinetics and competition with other non-enzyme targets.
For a deep understanding of a compounds’ molecular MOA, structural approaches can help elucidate not only mechanism of inhibition but how and where a compound interacts with a target protein. Sygnature Discovery exploits both NMR and Crystallography to characterise compound binding at a structural level. Sygnature Discovery’s scientists have an extensive repertoire of skills and techniques available that allow a thorough characterisation of Mechanism of Action:
- In depth characterisation of mechanism action can be carried using classical enzymology methods
- Tight binding and reversibility can be identified by kinetic and biophysical methods
- Biophysical assays allow characterisation of compound MOA on a full range of targets
- Detailed understanding of the compound binding site can be determined by structural biology
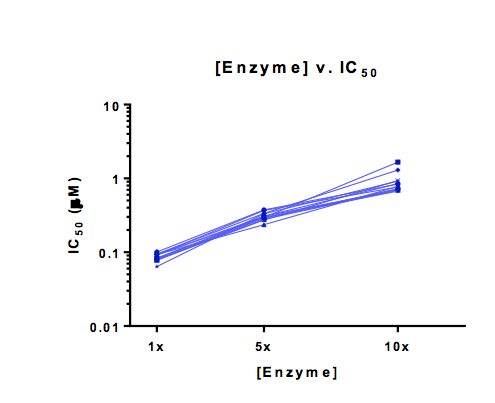
Figure A: [Enzyme] v. IC50 relationship is a way to determine if compounds have reached the tight binding limit of an assay i.e. when compounds are so potent they bind in a 1:1 manner and IC50 values can no longer be improved. This can be confirmed by increasing [enzyme] and looking for a concomitant increase in IC50 as shown above.
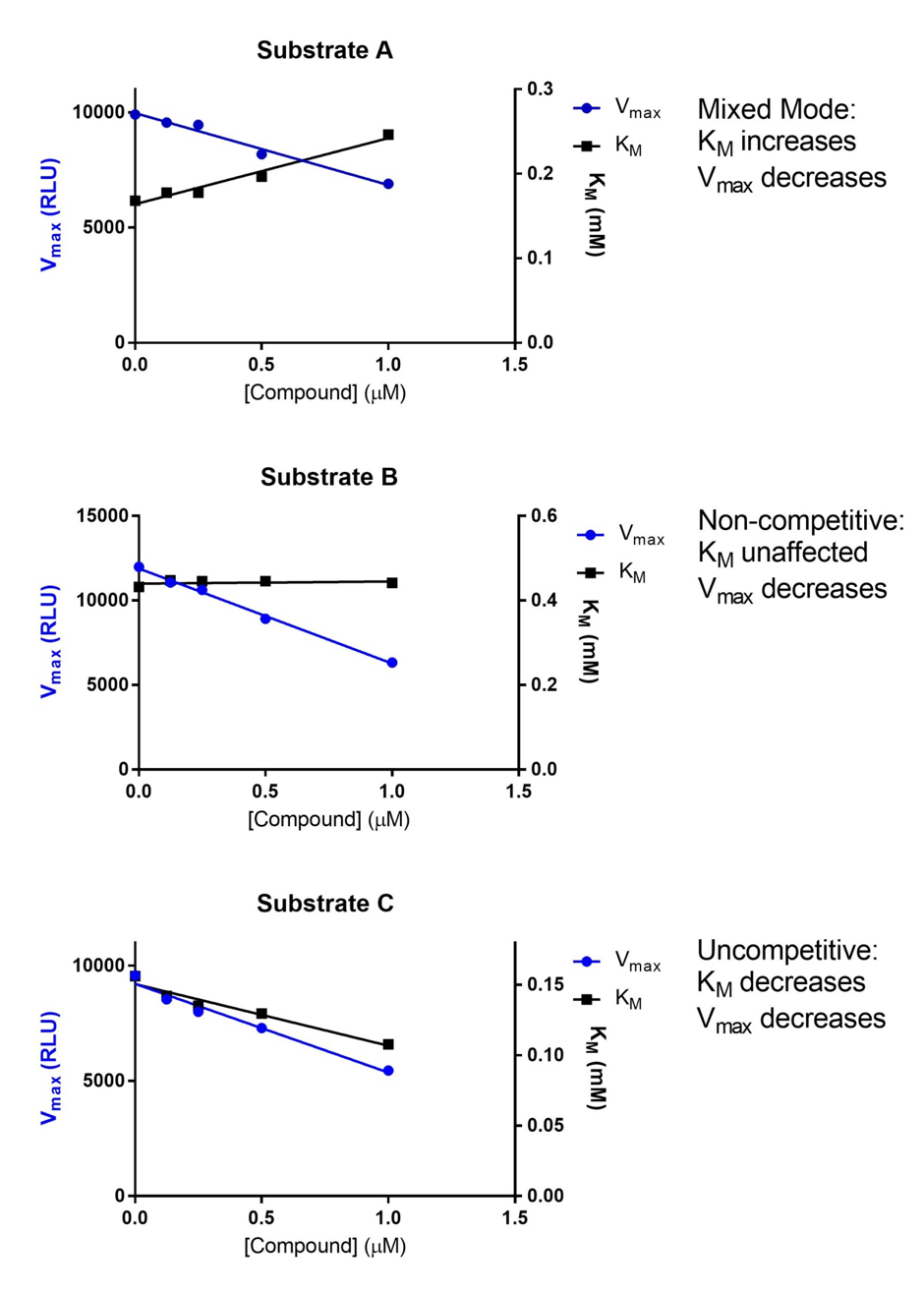
Figure B: For a complex, multi-substrate enzyme a single compound can show multiple mechanisms of action. The data above illustrates how compound concentration effects Michaelis-Menton kinetics and how that can be used to infer Mechanism action.
