HTS for drug discovery; how to enable successful hit finding
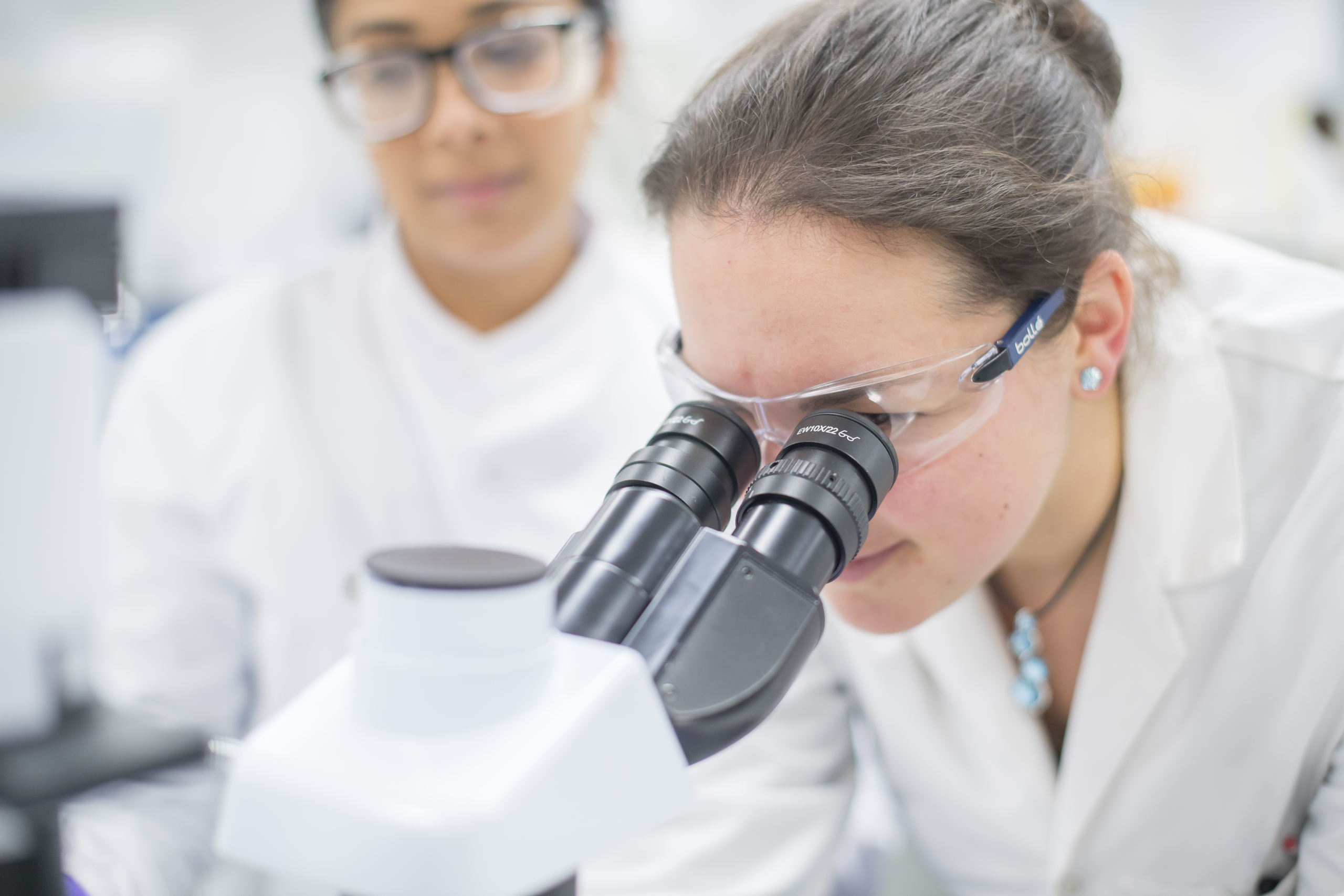
High throughput screening (HTS) is a well-established process for rapidly identifying novel chemical starting points for drug discovery programmes. It involves the seamless integration of several key scientific functions: medicinal chemistry, biology, informatics and advanced automation technology.

For HTS to succeed, the primary screen must use a high-quality, structurally diverse compound library, which represent attractive starting points for new drug discovery programmes. Ideally library compounds exhibit developable lead-like attributes, desirable physical chemistry properties, and no undesirable substructures that might cause problems further on in the discovery process.
The overall aim is to rapidly identify compounds with useful biological activities from a structurally diverse library. Ideally the biological assay itself must be disease-relevant and modified to make it amenable to HTS, which is achieved by optimising assay and automation parameters such as plate density, assay volume, long term reagent stability and tolerance of reagents to handling by bulk liquid dispensers.
There is far more to a successful HTS campaign than simply choosing the optimal primary screen. A well-planned post HTS assay cascade is critical to triage any potential HTS active compounds. This “Hit Evaluation” process can include a counter screen to identify and remove compounds that, for example, directly interfere with assay end point detection rather than the enzyme target per se. It can also be important to use an orthogonal assay to confirm compound target engagement, such as a biophysical assay to determine target binding affinity. Other components of the cascade can include target selectivity assays to check for selectivity against any related, undesirable targets, and a medicinal chemistry assessment.
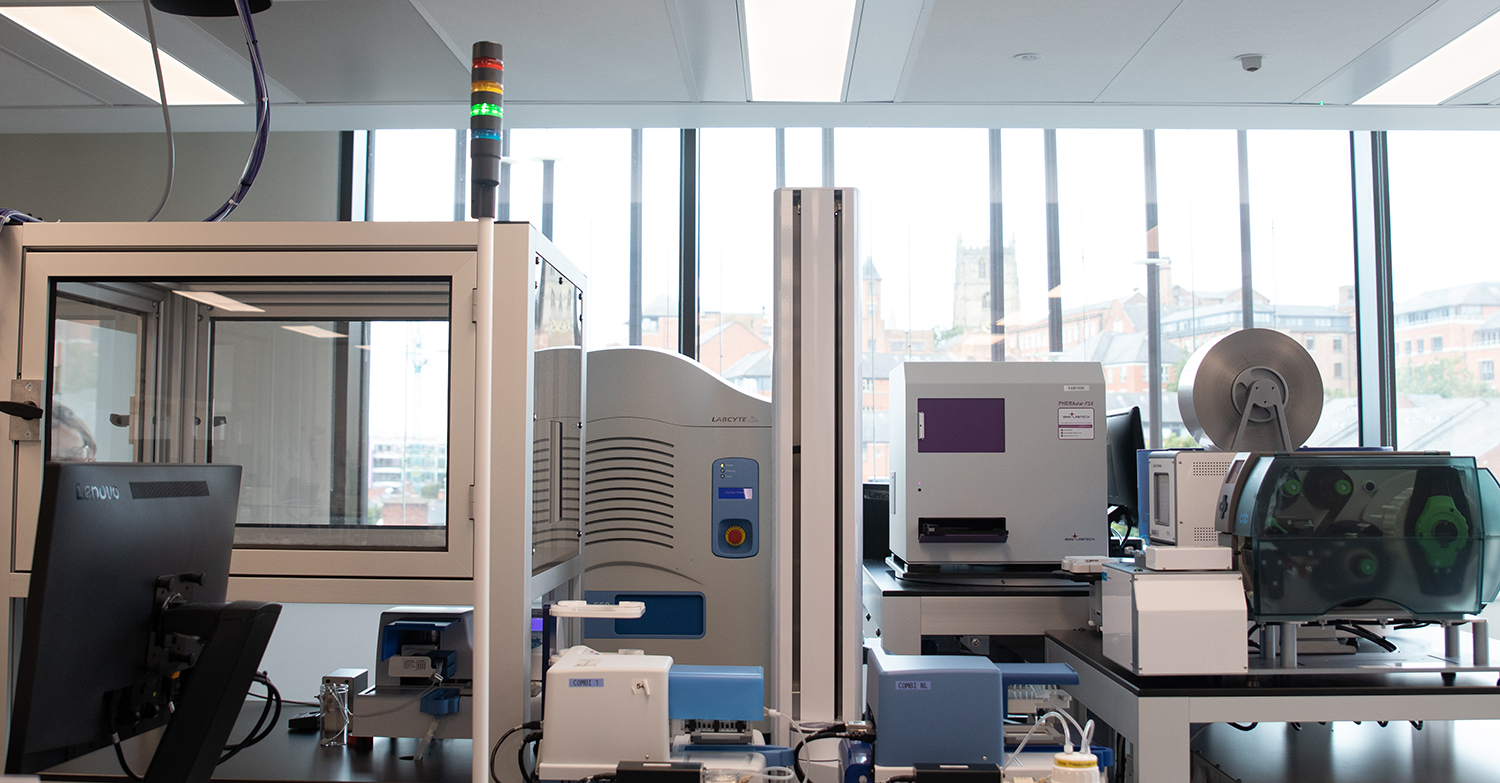

Any HTS system must integrate the right equipment. It should include a highly flexible automation platform that can accommodate different assay types and operate at scale cost efficiently. Preferably the automation is controlled by dynamic scheduling software, to enable flexibility in the type and construction of screen that can be undertaken, and coordinate in-time communication between the compound management system, the automation of the HTS itself, and data analysis functions.
HTS information technology systems must be able routinely handle, visualise and interrogate large complex assay data sets. If suitable informatics are not available, this can prevent the screening information from being used effectively such that potentially valuable performance meta information can easily be missed. Without a doubt, high-quality informatic systems are essential for maintaining the unified automation of the HTS process, and to the analysis and interpretation of large-scale screening data.
We took all of these important points into account when designing our HTS facility . As we started from scratch, we did not have to design our new HTS unit using existing legacy systems, but used industry-leading suppliers to design a HTS solution based around their leading-edge platforms. This together with our carefully selected and curated Lead Finder library will help to maximise the probability of identifying viable chemical hits for our clients’ programs.
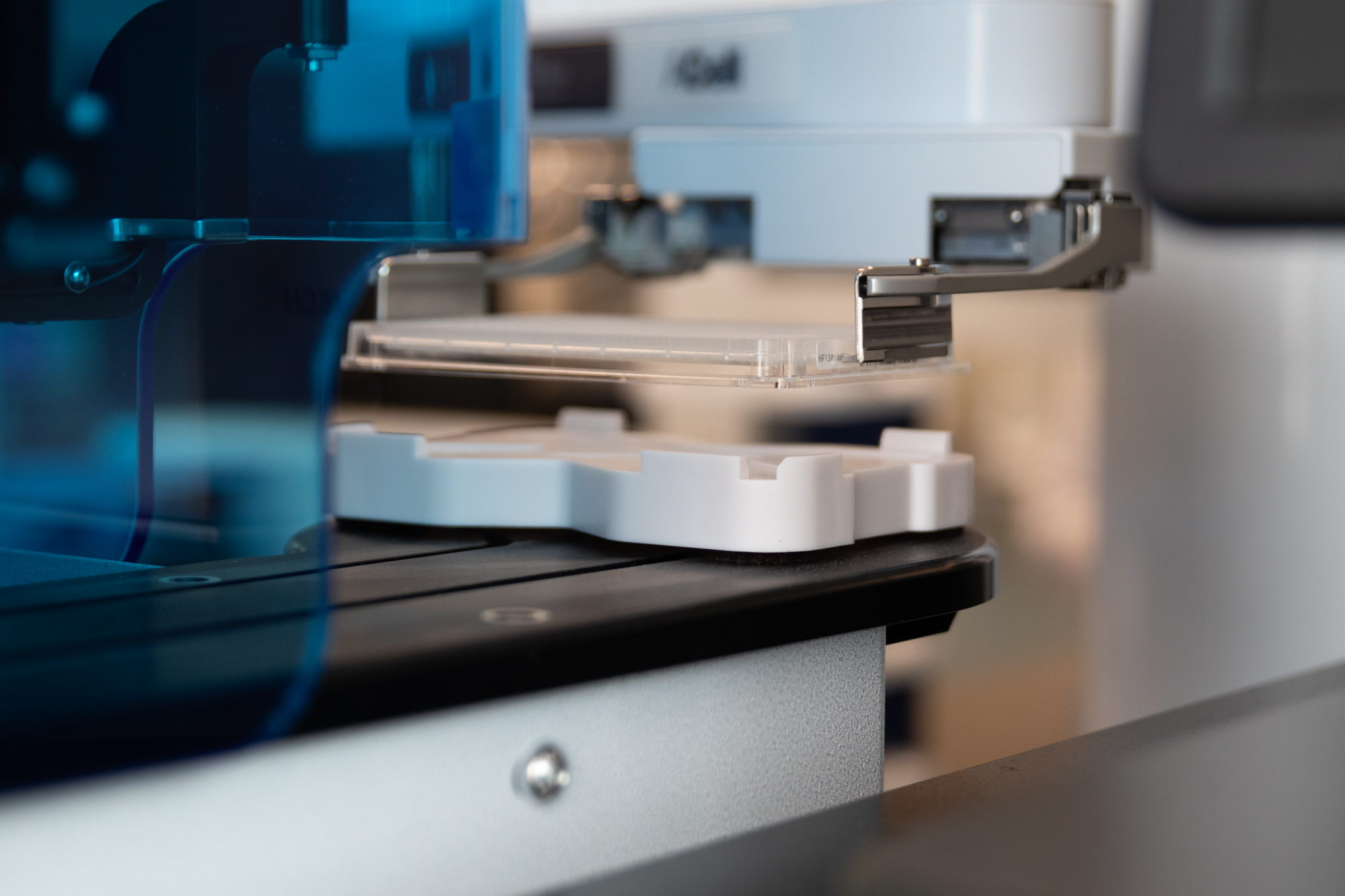

The Sygnature Discovery HTS system has a fully integrated informatics platform to ensure compound and assay data fidelity. We can design and order assay plates in Titian Mosaic sample management software, run the fully automated screening process on our HighRes automated system, and run data analysis and approval in Genedata Screener. This automated HTS workflow decreases the potential for human errors and removes potential bottlenecks, while maintaining cost efficient use of time and resources.
We hope you’ve found this article helpful; if you would like to discuss HTS, how we can help your project, or even drug discovery in general then we’d love to hear from you. You can get in touch by using any of the contact forms.