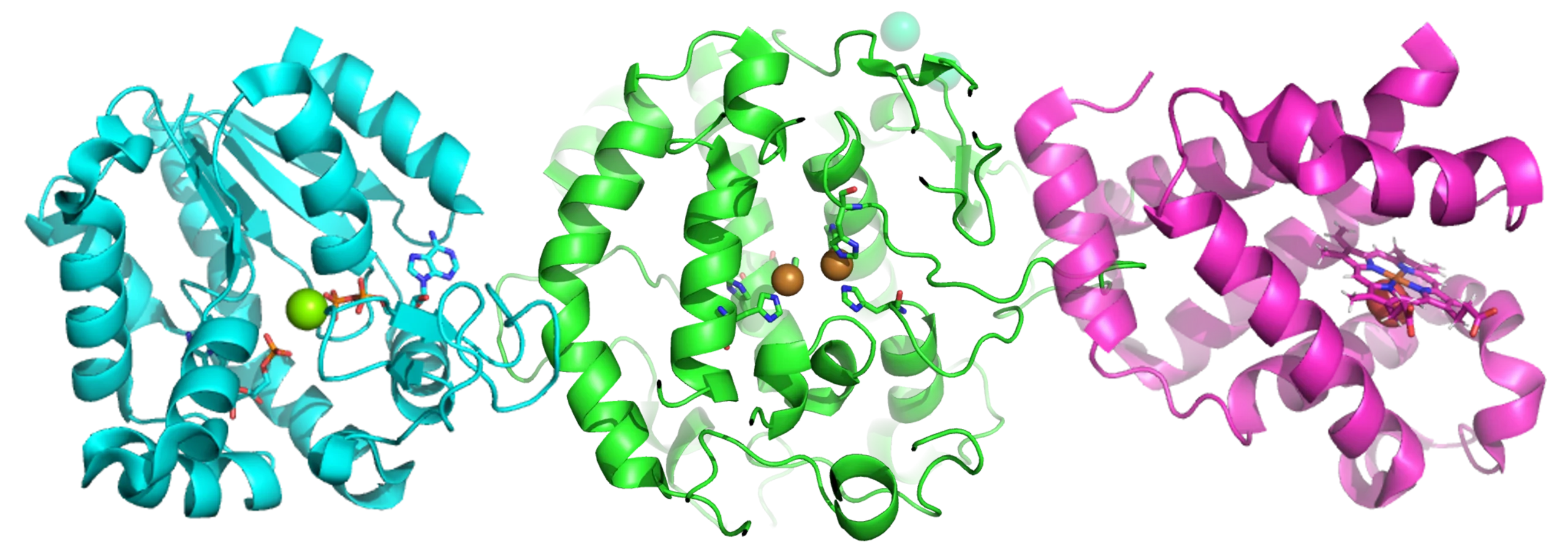
The revolutionary field of Ribonucleic acid (RNA) based drug discovery could lead to life-changing medicines in therapeutic areas such as cancer and infectious diseases. As well as novel targets on RNA-modifying enzymes, it encompasses drugs that interact with RNA sequences themselves.
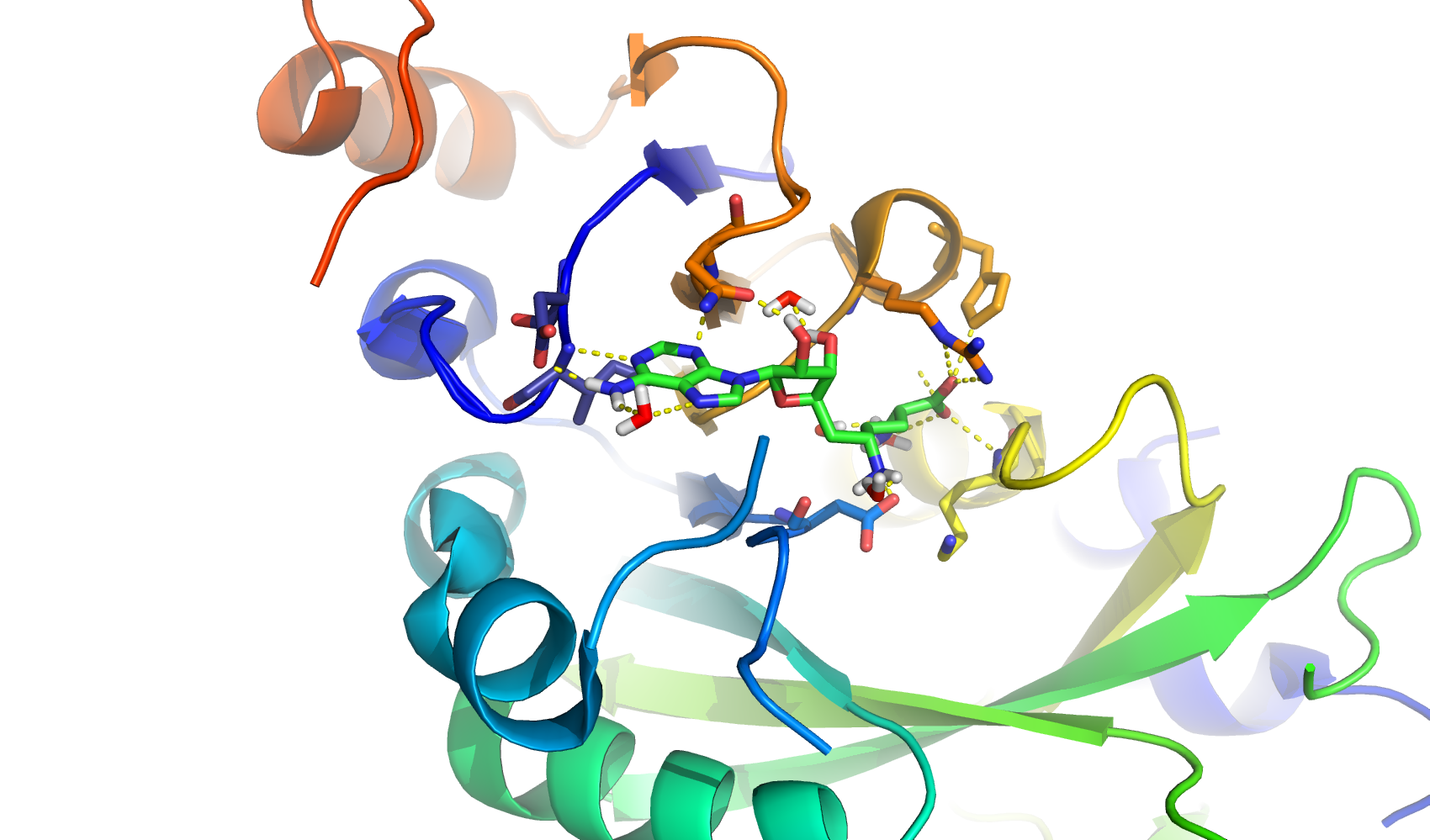 RNA-modifying enzymes include those that recognise, add or remove methyl groups from RNA bases such as adenosine. RNA sequence targets, meanwhile, include both tumour-suppressing microRNAs and their precursors. Drugs acting on any of these target pathways could affect gene transcription, with potential therapeutic benefits in cancer and other diseases.
RNA-modifying enzymes include those that recognise, add or remove methyl groups from RNA bases such as adenosine. RNA sequence targets, meanwhile, include both tumour-suppressing microRNAs and their precursors. Drugs acting on any of these target pathways could affect gene transcription, with potential therapeutic benefits in cancer and other diseases.
These new drugs are likely to look rather different from those that act at more traditional protein targets. So how do we go about finding hits, and where in chemical space will we find molecules that will interact with the targets? What selectivities are required, or desirable, if we are to design well-tolerated medicines?
New types of assay are required to screen libraries of pharmacologically active compounds for RNA targets, in order to identify hit molecules from them. Sygnature scientists have recently put a great deal of effort into developing assays and biophysical techniques that will underpin RNA-based drug discovery. These include:
Microscale thermophoresis (MST)
 Many different biophysical techniques offer huge potential in advancing the field. For example, we have developed several biophysical assay formats, notably using MST. Using this technique, we have created assays that assess potential inhibitors of enzymes that modify RNA by removing a methyl group from N-6-methyladenosine in the RNA; these enzymes are often referred to as erasers. An example of its application is on the RNA that makes the fat mass and obesity associated protein FTO, which has been revealed to regulate self-renewal in cancer cells and energy intake in obesity.
Many different biophysical techniques offer huge potential in advancing the field. For example, we have developed several biophysical assay formats, notably using MST. Using this technique, we have created assays that assess potential inhibitors of enzymes that modify RNA by removing a methyl group from N-6-methyladenosine in the RNA; these enzymes are often referred to as erasers. An example of its application is on the RNA that makes the fat mass and obesity associated protein FTO, which has been revealed to regulate self-renewal in cancer cells and energy intake in obesity.
Surface plasmon resonance (SPR)
 Biophysical assays using SPR, and enzyme assays using fluorescence-based readouts, have also been developed. These assays can be used to study binders of microRNA sequences that might inhibit tumour-suppressing RNAs. A notable example is the fluorescence-based assay we have designed for inhibitors of the Dicer enzyme-mediated processing of microRNAs from their precursors. These inhibitors may have significant effects on the growth and invasiveness of tumours, particularly in glioblastomas, a particular type of brain tumour that’s been notoriously hard to treat.
Biophysical assays using SPR, and enzyme assays using fluorescence-based readouts, have also been developed. These assays can be used to study binders of microRNA sequences that might inhibit tumour-suppressing RNAs. A notable example is the fluorescence-based assay we have designed for inhibitors of the Dicer enzyme-mediated processing of microRNAs from their precursors. These inhibitors may have significant effects on the growth and invasiveness of tumours, particularly in glioblastomas, a particular type of brain tumour that’s been notoriously hard to treat.
Mass spectrometry
 We have also developed assays using mass spectrometry to detect inhibitors of enzymes that add methyl groups to the N-6-methyladenosine in RNA, otherwise known as writers. A good example here are the methyltransferase-like 3/14 enzymes which are critically involved in the regulation of genes leading to unrestricted cancer growth.
We have also developed assays using mass spectrometry to detect inhibitors of enzymes that add methyl groups to the N-6-methyladenosine in RNA, otherwise known as writers. A good example here are the methyltransferase-like 3/14 enzymes which are critically involved in the regulation of genes leading to unrestricted cancer growth.
We have already presented several posters showcasing these techniques at a number of high-profile scientific conferences, including the American Association for Cancer Research (AACR), the European Laboratory Research & Innovation Group (ELRIG), and STORM Therapeutics meeting. These disclosures have highlighted our growing expertise in the field of RNA-modifying enzyme drug discovery, and further presentations in the RNA target area will follow shortly.
The potential that RNA-based drug discovery offers is truly exciting. We are already progressing projects involving RNA-modifying targets, and these targets offer huge potential for important advances in medicine, particularly in the field of oncology. We believe that the knowledge and capabilities that we are developing will enable success on these challenging, exciting targets, with the aim of delivering groundbreaking first-in-class therapeutics to patients.
We continually engage with our industry on a range of drug discovery topics. If you would like to discuss our bioscience capabilities or drug discovery in general then we’d love to hear from you. You can get in touch by using any of the contact forms.
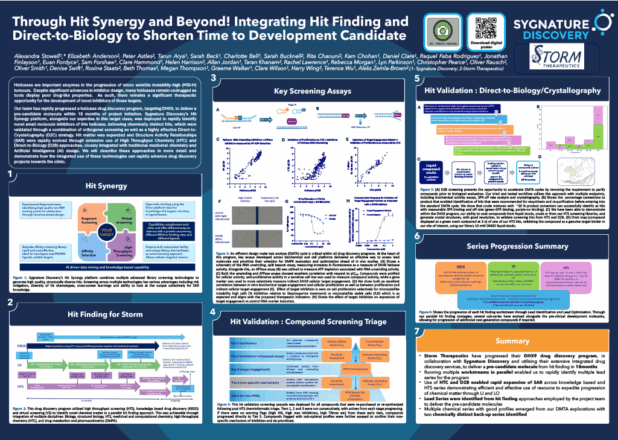
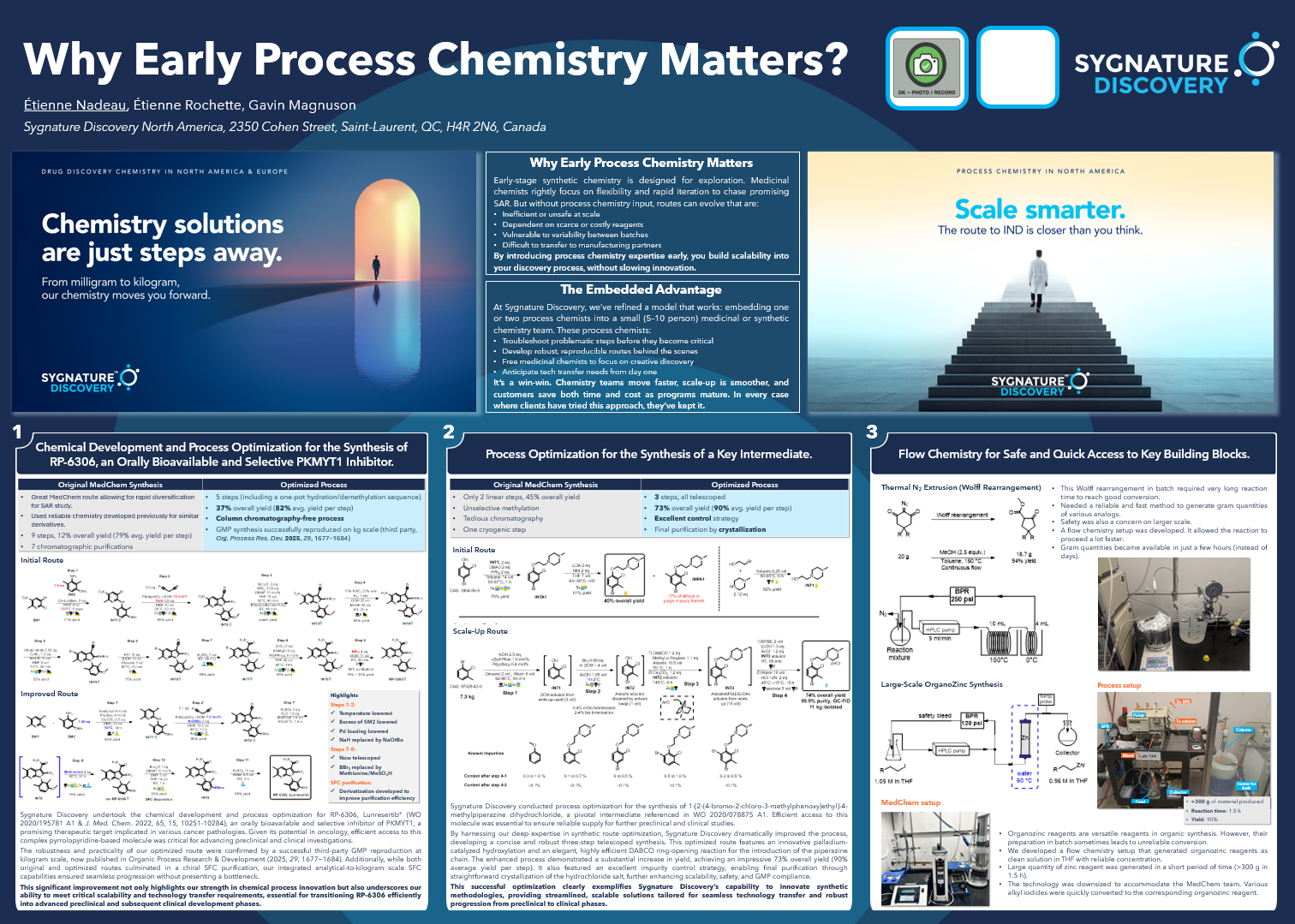



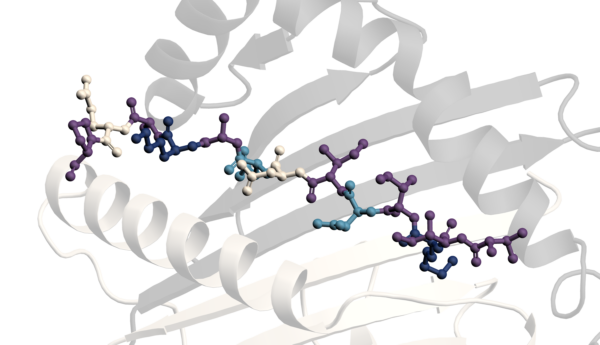











 Many different biophysical techniques offer huge potential in advancing the field. For example, we have developed several biophysical assay formats, notably using MST. Using this technique, we have created assays that assess potential inhibitors of enzymes that modify RNA by removing a methyl group from N-6-methyladenosine in the RNA; these enzymes are often referred to as erasers. An example of its application is on the RNA that makes the fat mass and obesity associated protein FTO, which has been revealed to regulate self-renewal in cancer cells and energy intake in obesity.
Many different biophysical techniques offer huge potential in advancing the field. For example, we have developed several biophysical assay formats, notably using MST. Using this technique, we have created assays that assess potential inhibitors of enzymes that modify RNA by removing a methyl group from N-6-methyladenosine in the RNA; these enzymes are often referred to as erasers. An example of its application is on the RNA that makes the fat mass and obesity associated protein FTO, which has been revealed to regulate self-renewal in cancer cells and energy intake in obesity. Biophysical assays using SPR, and enzyme assays using fluorescence-based readouts, have also been developed. These assays can be used to study binders of microRNA sequences that might inhibit tumour-suppressing RNAs. A notable example is the fluorescence-based assay we have designed for inhibitors of the Dicer enzyme-mediated processing of microRNAs from their precursors. These inhibitors may have significant effects on the growth and invasiveness of tumours, particularly in glioblastomas, a particular type of brain tumour that’s been notoriously hard to treat.
Biophysical assays using SPR, and enzyme assays using fluorescence-based readouts, have also been developed. These assays can be used to study binders of microRNA sequences that might inhibit tumour-suppressing RNAs. A notable example is the fluorescence-based assay we have designed for inhibitors of the Dicer enzyme-mediated processing of microRNAs from their precursors. These inhibitors may have significant effects on the growth and invasiveness of tumours, particularly in glioblastomas, a particular type of brain tumour that’s been notoriously hard to treat. We have also developed assays using mass spectrometry to detect inhibitors of enzymes that add methyl groups to the N-6-methyladenosine in RNA, otherwise known as writers. A good example here are the methyltransferase-like 3/14 enzymes which are critically involved in the regulation of genes leading to unrestricted cancer growth.
We have also developed assays using mass spectrometry to detect inhibitors of enzymes that add methyl groups to the N-6-methyladenosine in RNA, otherwise known as writers. A good example here are the methyltransferase-like 3/14 enzymes which are critically involved in the regulation of genes leading to unrestricted cancer growth.